29 HE切片细胞类型
HE切片的细胞类型鉴定
看到一篇2025-12-22日上线的预印版文献(1),报道了Classpose,可用于HE染色全片的细胞类型鉴定。


1. 两种细胞分割/cell segmentations
Instance segmentation … aims to determine whether a pixel belongs to a unique cell (1).
以HE切片为例,直观理解就是选切片上的细胞(核)。
Semantic segmentation … aims to determine to which cell type a pixel belongs to (1).
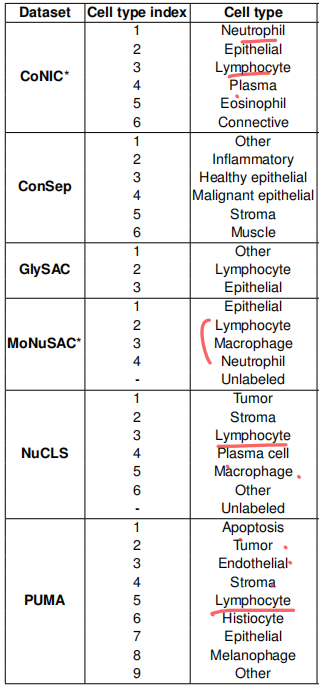
以HE切片为例,直观理解就是确定每个细胞(核)属于哪种类型(比如中性粒细胞、上皮细胞、淋巴细胞等等)。
2. Classpose的原理?
看不懂文献的方法,好像也没有读懂的动力。
3. Classpose能做什么?
Classpose, as easily trainable analog of the Cellpose-SAM model for semantic segmentation (1).
4. Classpose的优点?
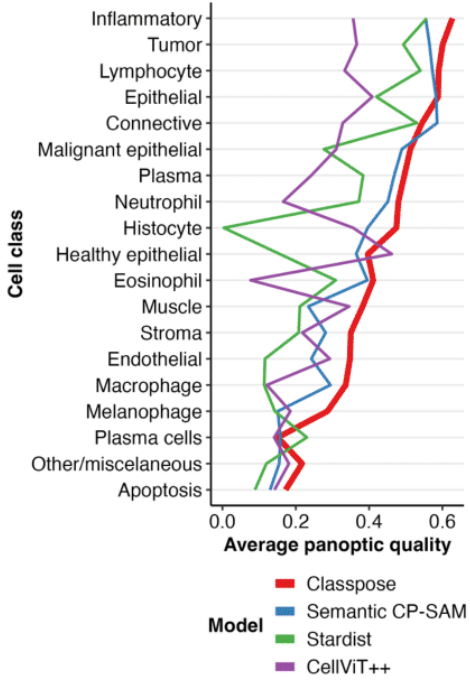
We extensively benchmark Classpose against 3 other state-of-the-art methods (Semantic Celllpose-SAM, CellViT++, and Stardist) across 6 datasets (CoNIC, ConSep, GlySAC, MoNuSAC, NuCLS, and PUMA), showing that Classpose consistently outperforms all other methods (1).
5. Classpose如何使用?
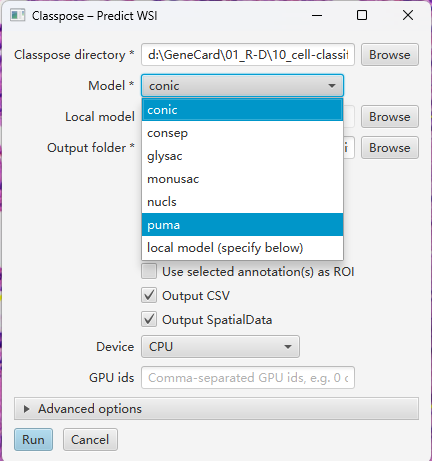
… provide a commond-line tool and a QuPath extension for whole slide-image … (1)
6. 自己安装Classpose的QuPath extension体验下
具体安装过程有些折腾,但最终还是成功了。安装过程参考(2) (3)。
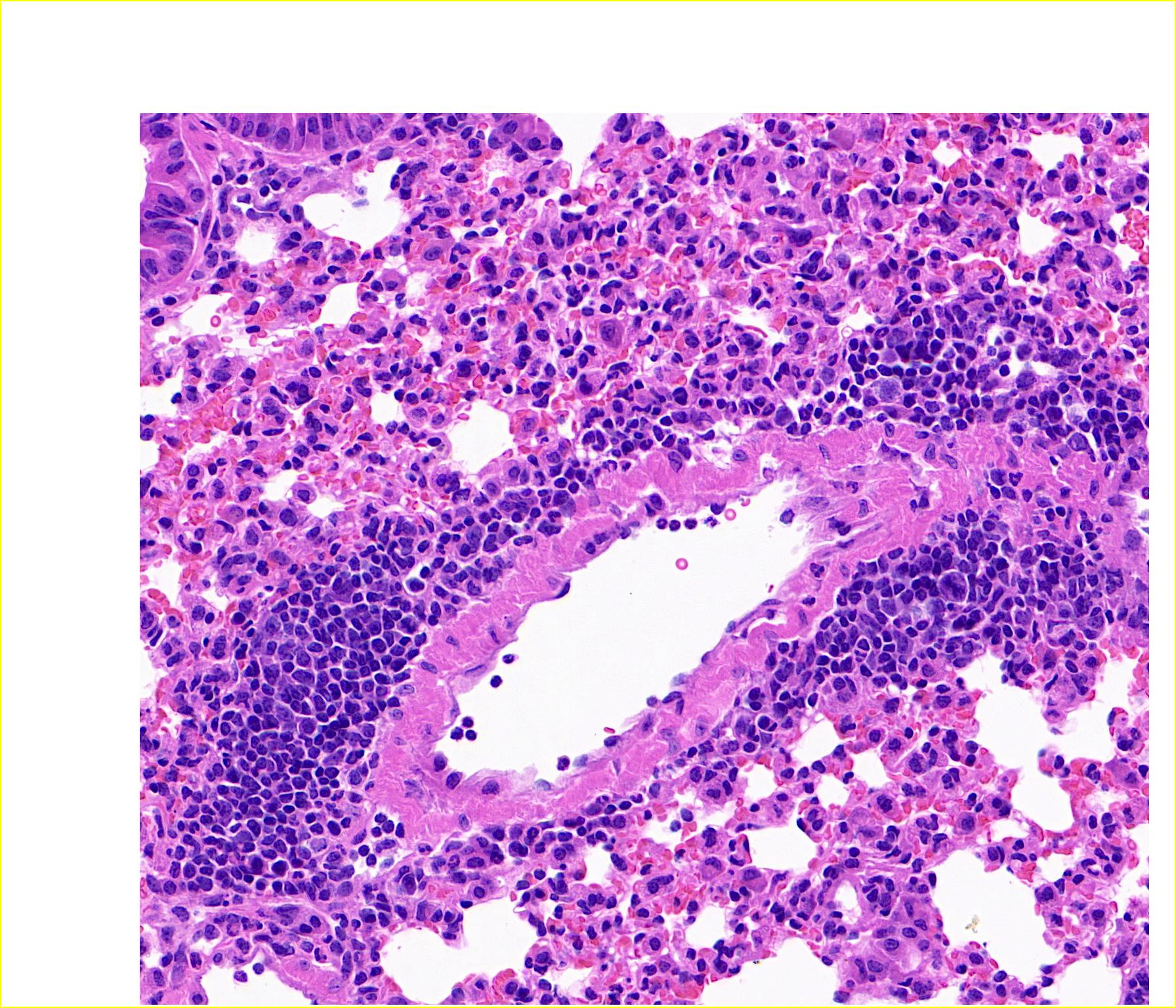
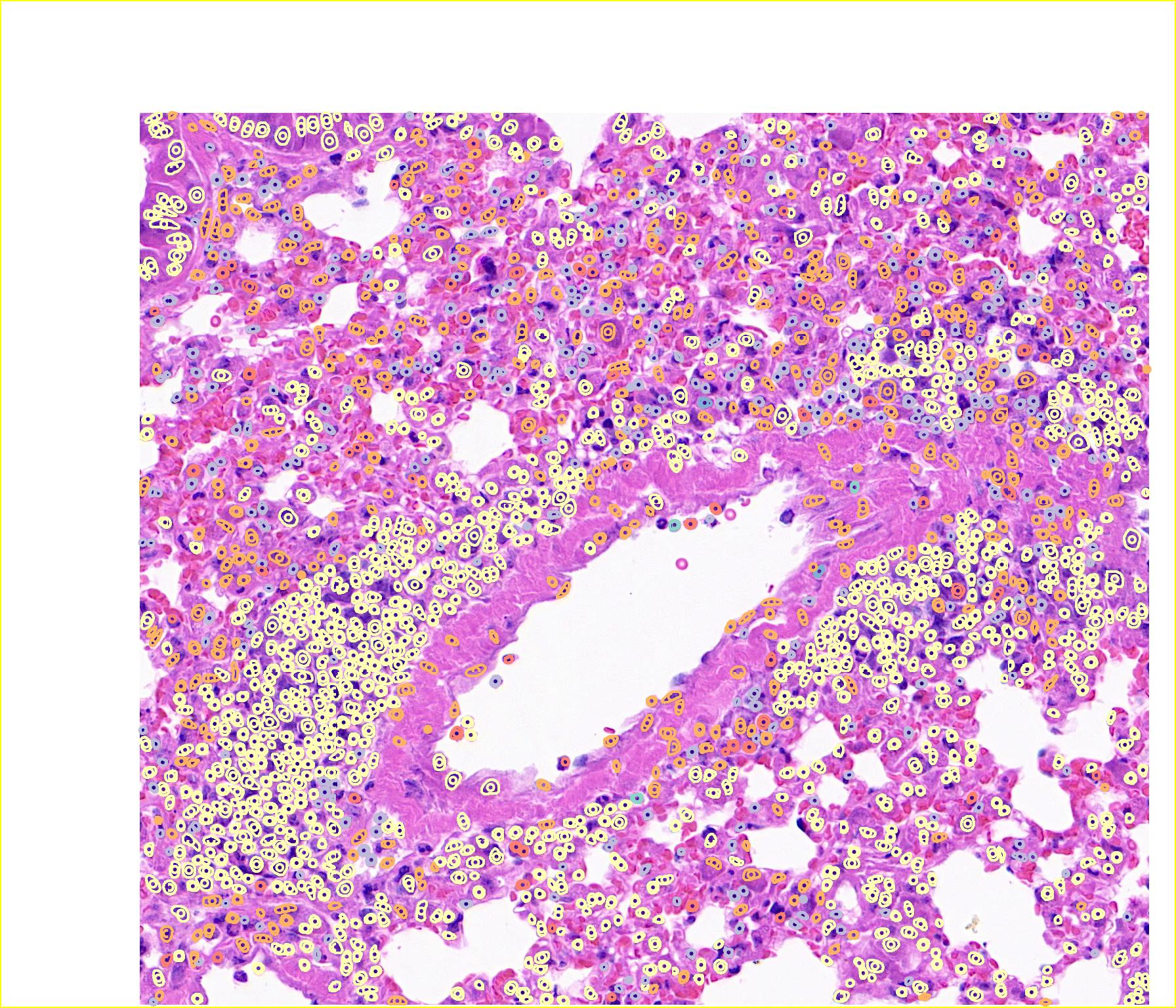
7. 期待Classpose文章正式发表
目前该软件工具还是预印版,期待正式在某个科研杂志发表,这样使用起来更加有底气些。
本着严谨的态度,感觉是不是应该比较下Classpose的预测结果和多色免疫荧光染色的结果一致性?
或者基于Classpose分析得到了某种结论,再针对该结论通过某种合适的实验验证下?